Multi-disciplinary Team is Unlocking the Connected Secrets of Neurons
Multi-disciplinary Team is Unlocking the Connected Secrets of Neurons
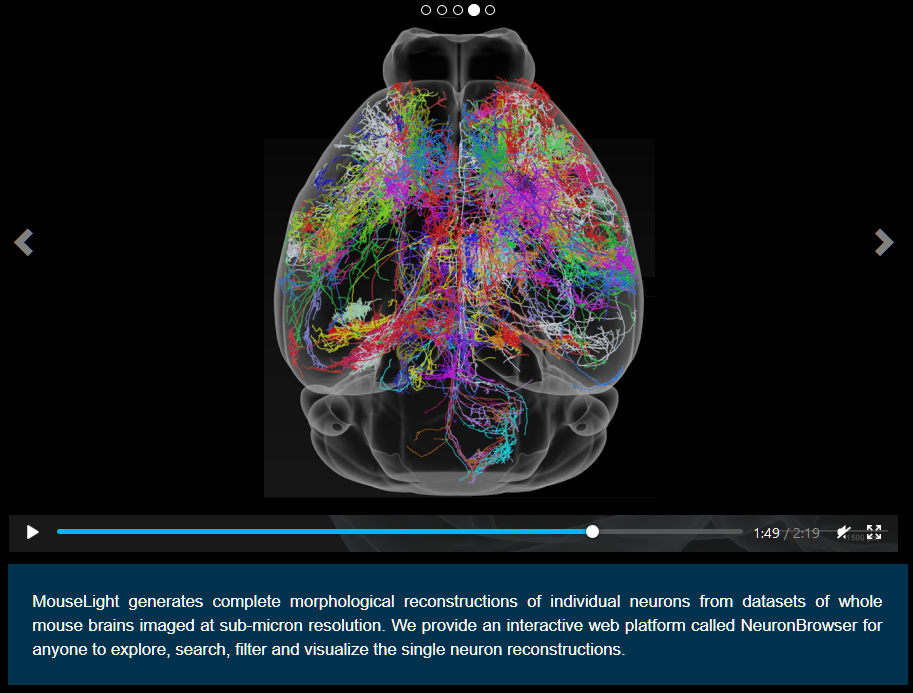
An amazing multi-disciplinary team of about 40 neurobiological and computer scientists and technicians is systematically cataloging, tracing and visualizing neurons in the mouse brain. The effort, called MouseLight, is funded by the Howard Hughes Medical Institute and is conducting the work at HHMI’s spectacular Janella Research Campus in Virginia. MouseLight is generating maps of individual neurons across the entire mouse brain using high-speed, high-resolution light microscopy. So far, 300 neurons have been painstakenly completed, with a target of 1000 by the end of 2018. While impressive, there is still a considerable ways to go to capture all of the estimated 70 million! neurons in the mouse brain.
The visualizations of the neurons, singly and in combination, are just amazing. For example, according to a recent project news release, some “axons of individual neurons in the thalamus often branch profusely in unexpected combinations of cortical areas, such as regions involved in taste, touch, and movement. Similarly, in the subiculum, a region involved in learning and memory, neurons almost always reach out to a few different places. In the neocortex, the six-layered structure associated with the highest cognitive functions, many single-neuron projections spread expansively. One neuron the researchers traced ran scattershot across the cerebral cortex, sending long, branching axons arcing across both hemispheres like a fireworks explosion.”
At the project’s Web site you can see an introductory video and use an interactive Neuron Browser, which allows you to select and view and manipulate specific neurons, including a nice tutorial and pre-canned examples:
Only one to a few neurons may be processed at a time to generate the primary data underlying all of this analysis. The basic process works as follows: First, the researchers inject the mouse brain with a virus that highlights those few neurons. The brain is then excised and ‘cleared’ to enable light to penetrate the tissue. The brain is hit with a pulse of light to illuminate the target neurons using ‘two-photon microscopy’ to get the sharp images needed to identify the specific neurons, and then sliced (‘microtomed’) into 200-micrometer thick layers with a vibrating razor. After digitizing the image on the slice, the process is repeated until the entire brain is characterized. The digitized, illuminated areas on each slice are then re-constructed by computer with trained annotators tracing and confirming the actual neuron trace. Each processing of a few neurons yields approximately 20 terabytes of data. Neurons on the combined datasets are then color-coded, and are processed by 3-D visualization software for animations and rotations. The software, as demonstrated by the online browser, enables specific neurons to be turned on or off for inspection and other 3-D manipulations. As the team has continued to learn, it has continued to speed up the process.
These kinds of advances are now giving us an unprecedented look at the physical structure and connectedness of neurons. For those of us doing knowledge representation and digital cognition work, and if we believe ultimately our work needs to be a reflection of physical reality in some form, we are apparently still a long, long ways away from being able to model animate brain functions. The degrees of connectivity suggested are staggering, and well short of current practice in any digital artifact. One of the next challenges for neuroscience is to try to discover physical patterns of connections that shed light on generalizations, for it looks like, based on these early physical findings, that the brain is an unbelievably jumbled mess of pathways and connections. Finding general keys will be needed to unlock the mysteries.